Greetings!
This summer I will travel to the US and attend to three conferences: Transvision 2007 in Chicago, the World Future Society annual meeting in Minneapolis and the Science Foo Camp in Mountain View, California. I will speak about OS Biomedical Research in a student meeting associated to TV07, as it seems I am too young and unexperienced to be allowed on the main conference. Anyway, I will try to spread the meme and make more visible the problems that we have here dealing with diseases very well known but that have no cure, because they are not profitable for the industry.
In the WFS meeting I won't deliver any presentation, but I hope to discuss some of my work with the Millennium Project with the MP staff that will be there.
In Mountain View I will be at the famous Googleplex for the Science Foo Camp, new kind of conference, where there is no previous schedule, the contents are determined by the interaction of people. I hope to talk there about my current work at the MP, the importance of future studies, proper science education and skepticism, but also about OS Biomedical Research and the current efforts going in that way like The Synaptic Leap and current research done at my lab, testing the efficiency of already modelled compunds against Chagas' Disease. It would be a lot of fun, and I will try to learn as much as I can.
Here is my abstract on OS Biomedical Research:
Open Source Biomedical Research and Computational Biology can improve drug design and help to fight neglected diseases.
Abstract:
Several hundred millions of people around the world are affected by neglected diseases. One of the challenges they encounter is that the development of treatments for these diseases is not profitable for the private sector, as most of the affected are among the poorest people in the world. These diseases tend to attack people in tropical regions of Africa, Asia and the Americas, often in a chronic way, disabling people for years and causing further poverty and decay, in a downward spiral. Until now, the process of developing new drugs has been cumbersome and expensive, yielding many ineffective or highly toxic products, due to an approach based on random testing of substances.
Now, with new methods in computational chemistry, it is possible to design new drugs in rational way that targets specific parts of a given virus, bacteria or parasite, while it bypasses the equivalent parts in humans. This approach might save a great amount of time and resources previously wasted on useless compounds or on effective compounds against useless targets, which, in turn, could reduce the cost of drug development. However, even with these new tools, most commercial partners are still not interested in developing drugs for poor markets.
A possible solution to this problem is the application of the “Open Source” approach to drug development. The Open Source (OS) approach in software has shown it is possible to create useful, reliable and efficient products through a voluntary, collaborative process. In biology the OS approach could be used for collaborative non-profit research, aiming to avoid the duplication of efforts and the release of patent-free compounds for neglected diseases in a relatively short period of time, thanks to the new computational methods and distributed efforts; moreover, as these methods become increasingly efficient and the available information about pathogens grows (regarding genomes, expression patterns, etc.), a OS collaboration could make the process of drug development even cheaper and more efficient, and even allow for the creation of "backup drugs" to tackle the problem of drug resistance before it appears.
Keywords : Systems Biology, Open Source Biomedical Research, Neglected Diseases.
I am really surprised I got invited to Scifoo, given the extremely high level of the attendees and that I still (yes, shame on me, but I chose to take computational physics too) am an undergrad. But this is a perfect example fof the reasons I created this blog: In this age you don't have to be wealthy or live in the developed world to help to develop new things, to be in contact, to get opportunities like this. Even ten years ago this would have been impossible even in my wildest dreams. This, my readers, is a praise to globalization. I only am sad that many people smart enough to get opportunities like this is lacking a proper education. I dream of the day that is corrected and we, all mankind, can use our brains and hands to solve our current problems.





 A really funny picture, with a standing Tim O'Reilly at Google and an "Server not found" error on the web browser.
A really funny picture, with a standing Tim O'Reilly at Google and an "Server not found" error on the web browser. This is the way that a self organized schedule board looks. Personally, I love it, and the overall result was a wild display of creativity. It worked very well, not surprisingly.
This is the way that a self organized schedule board looks. Personally, I love it, and the overall result was a wild display of creativity. It worked very well, not surprisingly. Some of the attendants.
Some of the attendants.
 People at Google work really hard it seems. Even when you are pissing you can improve your coding! Keep going that way, guys, your work makes our lives easier. And I should learn to use time on that way.
People at Google work really hard it seems. Even when you are pissing you can improve your coding! Keep going that way, guys, your work makes our lives easier. And I should learn to use time on that way. James Randi, our secular saint!
James Randi, our secular saint! Greg Bear and Kim Stanley Robinson. Soon(?) I will post about their presentations. KSR was already one of my favorite SF writers. Now I like his work even more. I better save my opinion about Bear for later...
Greg Bear and Kim Stanley Robinson. Soon(?) I will post about their presentations. KSR was already one of my favorite SF writers. Now I like his work even more. I better save my opinion about Bear for later...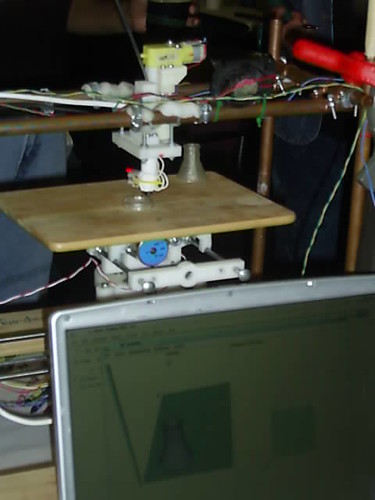 The RepRap, the macro almost-self-replicant machine that soon will manufacture its own pieces. A really mind blowing device, even in this early stage of development. If it can deliver its promises it will change a lot of things for a lot of people.
The RepRap, the macro almost-self-replicant machine that soon will manufacture its own pieces. A really mind blowing device, even in this early stage of development. If it can deliver its promises it will change a lot of things for a lot of people. Walking around that wonderful place, I stumbled upon the answer to the Ultimate Question!
Walking around that wonderful place, I stumbled upon the answer to the Ultimate Question! A truly Agalmic environment, according to me. Maybe I will write something serious about it when I am done with my thesis. Someday, I guess.
A truly Agalmic environment, according to me. Maybe I will write something serious about it when I am done with my thesis. Someday, I guess.




